HIV RNA quantification (or viral load) is a critical tool in the management of HIV disease. Detection of HIV RNA can help provide a positive diagnosis of HIV infection in certain clinical situations, such as acute or neonatal infection, where standard serological testing is inappropriate or unclear. Quantification of HIV RNA levels allows and predicts the rate of HIV disease progression[5] [6] and is the major laboratory tool for monitoring the response to antiretroviral therapy.
An increasing number of commercially available assays may be used as an aid in the diagnosis of HIV-1 infection, including acute or primary infection. The presence of HIV-1 RNA in plasma or serum of patients without antibodies to HIV-1 is indicative of acute or primary HIV-1 infection. Manufacturers are making claims to support the use of these assays as a supplemental test for specimens that have repeat reactive results with approved HIV immunoassays. There is a trend toward using approved tests to confirm HIV infection in serological test where antibodies are detected.
Nucleic acid quantification
Different technologies are used to quantitate HIV RNA but all have now achieved a sensitivity of 20-50 copies of HIV RNA per millilitre (mL) of plasma. In acute HIV infection, plasma HIV RNA levels may be used to assist in the confirmation of acute or primary infection before the appearance of HIV antibodies detected by an EIA or Western Blot assay. Most true-positive results will show very high levels (10^5 to 10^6 copies/mL) of circulating HIV RNA, consistent with uncontrolled viraemia. False-positive results occur in less than 10% of cases; these usually involve low RNA levels (less than 104 copies/mL) and are usually not reproducible[7] [8]. A number of manufacturers of HIV viral load tests still do not make claims that support the use of these tests for diagnosis of HIV infection.
Real time PCR assay
Conventional nucleic acid-based quantification methods are being replaced in some laboratories with one of a number of real time PCR assays that have become available. The Roche COBASTM AmpliPrep/ COBAS TaqManTM HIV-1 test combines automated sample preparation for HIV-1 RNA purification using the AmpliPrep, and real-time PCR amplification and detection using the COBAS AmpliPrep TaqManTMsystem. Similar to the Roche COBASTM Amplicor test, this assay targets dual conserved regions in HIV-1 and uses fluorescently-labelled probes to detect the amplified products in real time. The advantages are a wider dynamic range and greater sensitivity[9] [10]. Earlier versions of HIV viral load assays using single target amplification test systems were prone to variability in the levels of HIV-1 RNA reported to be caused by genetic diversity of HIV-1 and mutations and mismatches in the primer and probe target areas. The introduction of dual targets in highly conserved regions of the HIV-1 genome has reduced this variability and allows for greater equivalence in reported RNA results from patients with HIV infection with different subtypes.
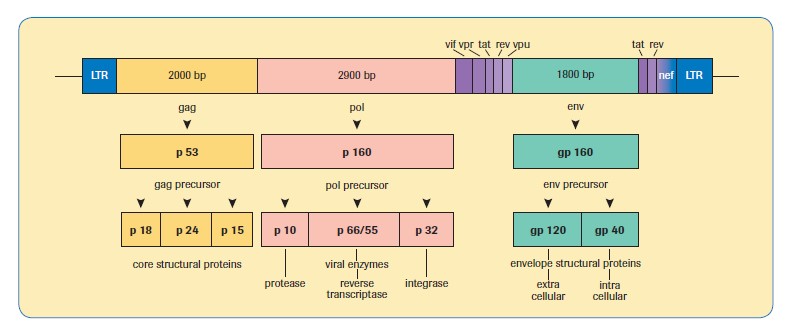
A comparison of different HIV-1 viral load methods is shown in Table 1, along with a diagrammatic representation of the genomic structure of HIV-1, for information, in Figure 3.
Table 1. Comparison of four commercial HIV-1 viral load methods
|
HIV Viral Load Method |
Assay Target/s |
Measuring Range (cp/mL) |
|
Roche |
gag and LTR |
20 - 1 X 107 |
|
Abbot |
Integrase |
40 - 1 X 107 |
|
Qiagen |
LTR |
34 - 4.5X107 |
|
Hologic |
pol and LTR |
30 - 1 X 107 |
Source: NSW State Reference Laboratory for HIV, St Vincent’s Hospital Sydney Limited.
Figure 3. Genomic structure of HIV-1
The Abbott RealTime HIV-1 assay, for use on the m2000 system, uses a unique partially double-stranded probe that targets the HIV-1 pol. The probe strands are labelled with a fluorophore (reporter) at the 5’ end, and a quencher moiety at the 3’ end of the shorter, complementary strand. In the presence of target pol sequences, the reporter probe preferentially binds, and upon release of the shorter quencher probe, fluoresces. Assay performance characteristics from Abbott Laboratories show that the assay has a 5-log10 linear range, assay specificity of 100% (n=259), 95% probability of detecting samples with a viral load of 25 copies/mL, and recognition of subtype panels from group M (A-H), group O, and group N.[11] [12] [13] [14] [15] [16]
The Hologic Aptima HIV-1 Quant Dx assay (Aptima assay; Hologic Inc., San Diego, CA) employs transcription-mediated amplification (TMA) and real-time detection for rapid, high-throughput testing of viral load on the Panther platform. Target amplification via TMA, utilizes two enzymes, MMLV (Moloney murine leukemia virus) reverse transcriptase and T7 RNA polymerase to amplify two regions of HIV-1 RNA (pol and LTR). Detection is achieved using single-stranded nucleic acid torches that are present during the amplification of the target and that hybridize specifically to the amplicon in real-time. Each torch has a fluorophore and a quencher. When the torch is not hybridized to the amplicon, the quencher is in close proximity of the fluorophore and suppresses the fluorescence. When the torch binds to the amplicon, the quencher is moved farther away from the fluorophore and it will emit a signal at a specific wavelength when excited by a light source. As more torches hybridize to an amplicon a higher fluorescent signal is generated. Assay performance characteristics from Hologic show that the assay has a 6-log10 linear range, assay specificity of 100% (n=630), 95% probability of detecting samples with a viral load of 30 copies/mL, and recognition of subtype panels from HIV-1 groups M, N and O. (38)
HIV DNA PCR
The HIV DNA assay detects both unintegrated and integrated forms of HIV DNA present in circulating peripheral blood mononuclear cells. There are a number of newly emerging qualitative assays able to detect both HIV DNA and RNA (total nucleic acid) for supplementary reference testing and diagnostic testing strategies. The main clinical use of these assays is in the qualitative detection of HIV in situations where serological testing is inappropriate (e.g. pre-seroconversion, or neonatal infection where results may be confounded by the presence of maternal antibodies) or where serological testing has been disputed or is inconclusive (e.g. indeterminate Western Blot assay result). The HIV DNA PCR test is highly sensitive (greater than 99%, detecting one copy of HIV DNA per 10,000 to 100,000 cells) and specific (98%), but has not replaced serology either as a screening test or as a diagnostic test in isolation. HIV DNA testing is the preferred test for early infant diagnosis when neonates are born to HIV seropositive mothers.
HIV DNA PCR and HIV RNA reverse transcriptase (RT)-PCR techniques were commonly used in laboratory research, however they have made a rapid transition to routine clinical practice. Amplification may be performed on various genomic regions of interest, and the products used in such analyses as HIV subtype determination, phylogenetic analysis, detection of genetic mutations, and prediction of viral tropism and drug resistance.[17] [18]
HIV antigen testing
Assays are available to directly detect the major structural core protein HIV-1 p24 (designated p24 because the molecular weight of the protein is 24 kilodaltons). While initially examined for utility as an assay to monitor HIV viral load[19] [20], it is no longer considered to be appropriate for viral load testing due to a lack of sensitivity[21] [22]. The p24 assay is widely used by reference laboratories in Australia, but is not commercially available in the USA and as such does not appear in recommended CDC testing algorithms. HIV-1 p24 antigen is particularly useful in diagnosis of acute HIV infection as a supplemental test when antibody tests may be negative, inconclusive or indeterminate. While nucleic acid tests offer superior sensitivity and specificity for the earliest detection of HIV infection, dedicated specimens are required for nucleic acid tests while HIV-p24 can still be performed on the primary serum sample submitted for antibody screening. The presence of HIV-1 p24 in newly identified sera or in sera which exhibit negative or inconclusive HIV-1 antibody tests should be confirmed by neutralisation with anti-p24 antisera.
In 2005, the so-called fourth generation HIV screening tests were developed. These tests detect HIV-1 p24 antigen and both HIV-1 and HIV-2 antibodies in a single test. They have been rapidly adopted globally as preferred screening tests for HIV infection and allow for the routine detection of recently acquired infections where antibody-only tests may be unreliable.

